Broadcasting Platform
Broadcasting link: All lectures will be streamed free of charge via Crowdcast and you can already register for the event under the following URL: https://www.crowdcast.io/e/prni-summerschool-2020/1
Attendee Quick Reference Guide for Crowdcast: Before you enter the event, please have a look at the instrucitons for attendees (click here)!
Preliminary schedule (CEST)
- Please be aware that this is a preliminary schedule, we hence reserve the right to alter the programs and speakers.
- Theoretically, the first two hours for each topic shown in the program are scheduled for the lecture, followed by two hours resereved for the tutorial. While depending on the actual content of each lecture, the duration of the lecture and the tutorial may slightly vary.
Keynote speakers
Peter Dayan
MPI for Biological Cybernetics, Germany

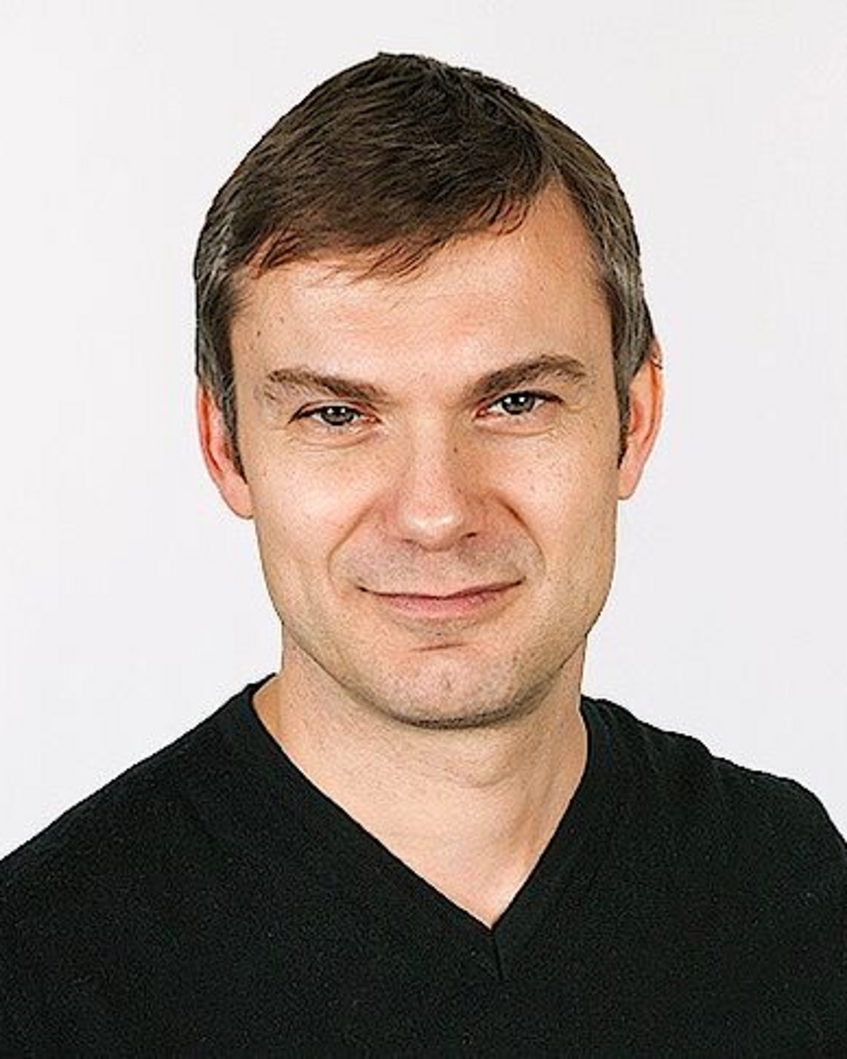
Biography: Peter Dayan studied mathematics at Cambridge University and received his doctorate from the University of Edinburgh. After postdoctoral research at the Salk Institute and the University of Toronto he moved to the Massachusetts Institute of Technology (MIT) in Boston as assistant professor in 1995. In 1998, he moved to London to help co-found the Gatsby Computational Neuroscience Unit, which became one of the best-known institutions for research in theoretical neuroscience, and was its Director from 2002 to 2017. He was also Deputy Director of the Max Planck/UCL Center for Computational Psychiatry and Ageing Research. In 2018, he moved to Tübingen to become a Director of the Max Planck Institute for Biological Cybernetics.
Lecture: Replay in Offline Planning
Abstract: Animals and humans replay neural patterns encoding trajectories through their environment, both whilst they solve decision-making tasks and during rest. There is also evidence that activity in sensory cortices is regenerated during periods of time without behaviour in a way that resembles its form when animals are actively engaged in perception. Under a common assumption that we build models of the world and recognize and plan actions using those models, such intrinsically generated patterns are ideal for various forms of model inversion, giving us access to fast and effective methods for sensory processing and decision-making. I will discuss our recent investigations using magnetoencephalography to detect replay in human subjects as they perform decision-making tasks. In a simple choice task, we found evidence for various forms of replay, which differed between subjects who flexibly adjusted their choices to changes in temporal, spatial and reward structure and those who were slower to adapt to change. The former group predominantly replayed comparatively less good trajectories during task performance, and subsequently avoided these inefficient choices. The latter replayed comparatively preferred, but suboptimal, trajectories during rest periods between task epochs. We suggest that online and offline replay both contribute to planning, but each are associated with distinct model-based and model-free decision strategies. Parts of this are joint work with Eran Eldar, Zeb Kurth-Nelson and Ray Dolan.
Katharina Dobs
MIT, USA

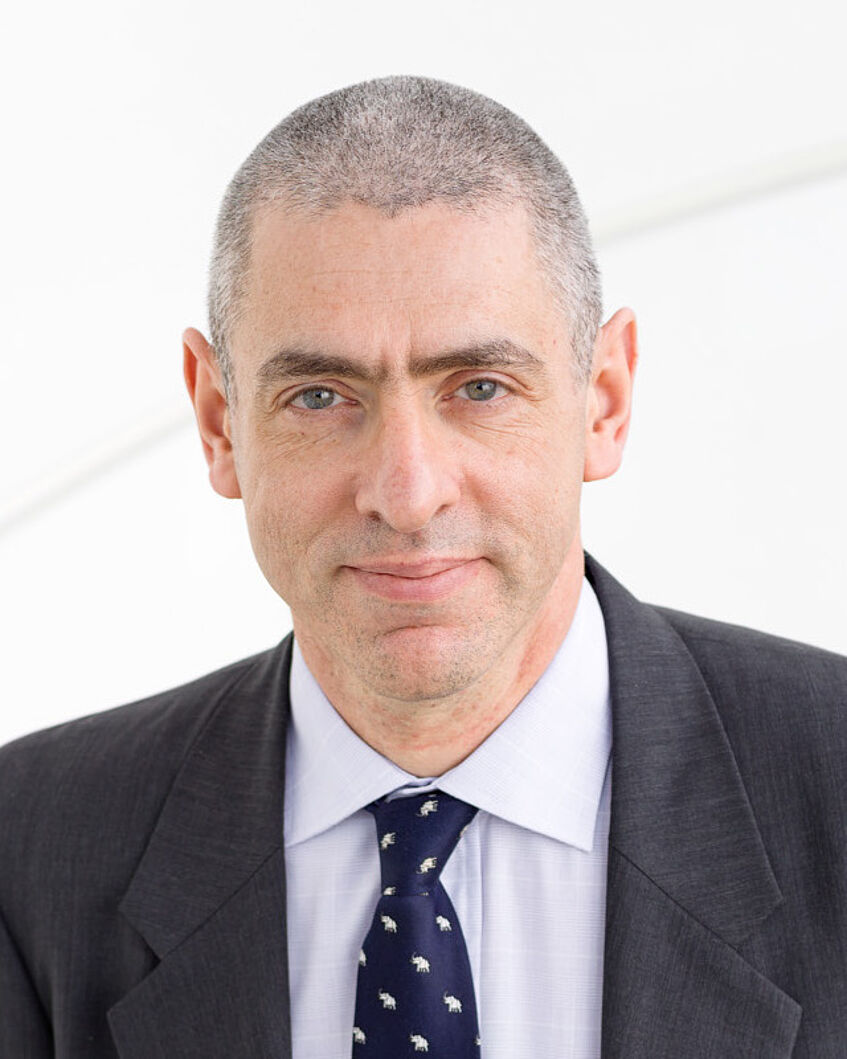
Biography: I am currently a postdoctoral researcher at MIT where I work with Nancy Kanwisher. I completed my PhD at the Max Planck Institute for Biological Cybernetics under the supervision of Isabelle Bülthoff and Johannes Schultz, investigating behavioral and neural correlates of dynamic face perception. During my first Postdoc at CNRS-CerCo working with Leila Reddy and Weiji Ma, I used a combination of Bayesian modelling, psychophysics and neuroimaging to characterize the integration of facial form and motion information during face perception.
Lecture: Using deep neural networks to understand why we have functional specialization in the human visual system
Abstract: Category-selective regions are a prominent feature of the ventral visual pathway. Why does the brain have specialization for some categories (e.g., faces, scenes), but not others (e.g., food, cars)? And why does functional specialization arise in the first place? In the first part of my talk, I use deep convolutional neural networks (CNNs) to test the hypothesis that face-specific regions are segregated from object regions because face and object recognition require different representations and computations. I show that a dual-task network trained on face and object recognition spontaneously segregates both tasks in the network. Critically, a dual-task network optimized for food and object categorization showed less task segregation. In the second part of my talk, I ask whether the representations learned by the dual-task network are predictive for activation in brain areas specialized for the same tasks. Specifically, I relate activation patterns extracted from the dual-task CNN to brain areas specialized for face and object processing measured using functional MRI. The results suggest a computational account of the observed specialization for faces in the human visual system, and demonstrate a general method for asking why, from a computational point of view, the brain is organized the way it is.
Romy Lorenz
University of Cambridge, UK& Stanford University, USA& Max Planck Institute for Human Cognitive and Brain Sciences, Germany

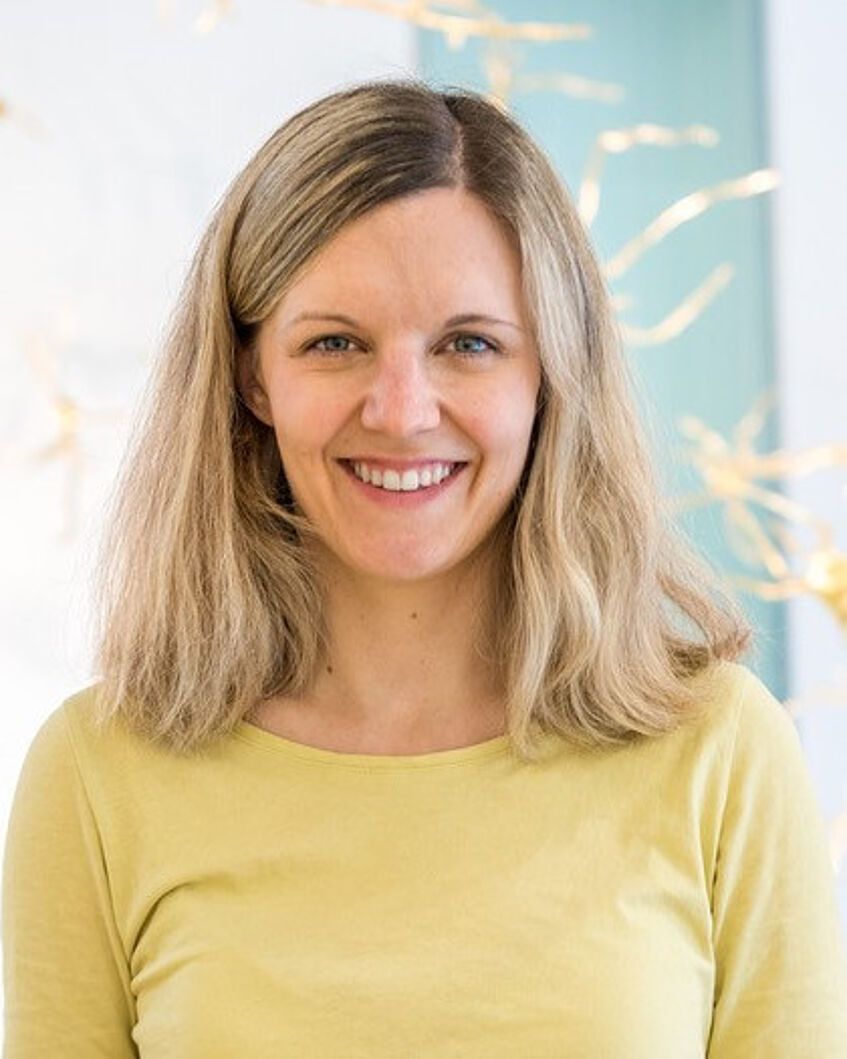
Biography: I am a Sir Henry Wellcome Postdoctoral Fellow at the University of Cambridge, Stanford University and the Max Planck Institute for Human Cognitive and Brain Sciences. My research interest lies in developing brain-computer interfaces (BCIs) and artificial intelligence (AI) with the aim to address cognitive neuroscience questions that have been historically challenging to tackle with conventional methods. By leveraging technological innovations, my long-term research vision lies in revisiting the classic taxonomy of cognitive processes and bring forward a neurobiologically-derived cognitive taxonomy. In 2017, I have completed my PhD in Neurotechnology under supervision of Robert Leech at Imperial College London. During my PhD, I have developed the “AI Neuroscientist”: a fMRI-based BCI that turns on its head how conventional cognitive neuroimaging experiments are performed. Before that, I completed my Masters in Human Factors at the Technical University Berlin. For this, I focused on EEG-based BCIs and gained research experience in labs in Berlin, Beijing and San Diego. Alongside this stream of research, I am interested in altered states of consciousness and passionate about novel ways of bringing together neuroscience and the arts.
Lecture: Neuroadaptive technology for cognitive neuroscientists
Abstract: Cognitive neuroscientists are often interested in broad research questions, yet use overly narrow experimental designs by considering only a small subset of possible experimental conditions. This limits the generalizability and reproducibility of many research findings. In this talk, I present an alternative approach that resolves these problems by combining real-time fMRI with a branch of machine learning, Bayesian optimization. Neuroadaptive Bayesian optimization is an active sampling approach that allows to intelligently search through large experiment spaces with the aim to optimize an unknown objective function. It thus provides a powerful strategy to efficiently explore many more experimental conditions than is currently possible with standard neuroimaging methodology. Alongside methodological details on non-parametric Bayesian optimization using Gaussian process regression, I will present results from three different studies where we applied the method to: (1) better understand the functional role of frontoparietal networks, (2) map cognitive dysfunction in aphasic stroke patients, and (3) tailor non-invasive brain stimulation parameters to a particular research questions. I will conclude my talk in discussing how Bayesian optimization can be combined with study preregistration to cover exploration, mitigating researcher bias more broadly and improving reproducibility.
Teaching faculty
Tonio Ball
University of Freiburg, Germany

Biography: Tonio Ball is head of the Neuromedical AI Lab at the University of Freiburg, Germany, where he also earned his MD. He works with intracranial and non-invasive EEG, including optimization of EEG acquisition, implant development, and application in BCIs, e.g., for the closed-loop online control of robotic systems. Recently, he focusses on application of deep learning techniques for improving online and offline decoding of information from EEG signals. He worked as a PostDoc for the Heidelberg Academy of Sciences and Humanities’s, and is a founding member of the Bernstein Center Freiburg and of the Cluster of Excellence BrainLinks-BrainTools in Freiburg.
Lecture: Deep Learning in neuroscience with a focus on EEG
Abstract: In this course you will learn how deep learning can be applied on EEG for classification and regression analysis; in addition advanced topics such as invertible networks for non-linear component analysis will also be touched. The course will start with a conceptual introduction and also include hands-on sessions using the braindecode open source package: https://braindecode.org/
Programming will be done in Python using Jupyter Notebooks, but no strong coding skills in Python are required.
Alexandre Gramfort
INRIA, France

Biography: Alexandre Gramfort is a senior researcher at INRIA, France, and formerly Assistant Professor at Telecom ParisTech, Université Paris-Saclay, in the image and signal processing department from 2012 to 2017. He is also affiliated with the Neurospin imaging center at CEA Saclay. His field of expertise is signal and image processing, statistical machine learning and scientific computing applied primarily to functional brain imaging data (EEG, MEG, fMRI). His work is strongly interdisciplinary at the interface with physics, computer science, software engineering and neuroscience. He has coauthored more than 30 journal papers and 50 conference papers since 2009 and has received a number of awards since the beginning of his research career: EADS PhD award in the category interdisciplinary research, young investigator awards at the international conference Biomag in 2010, and international conference HBM in 2011, runner up Erbsmann's award in 2015 at the IPMI conference. Alexandre Gramfort is committed to open source software development. He is a core developer of the Scikit-Learn machine learning software (http://scikit-learn.org) which is heavily used both in industry and in academic research. He is at the origin and the leader of the development of the MNE-Python software (https://mne.tools) now used and developed across many labs worldwide. In 2015, he was awarded a Starting Grant by the European Research Council (ERC).
Lecture: Hands on machine learning on MEG/EEG data with the MNE software
Abstract: In this course you will learn how machine learning can be applied on MEG/EEG to predict from evoked or induced brain activity. The course will balance theoretical presentations of the algorithms and hands sessions using the MNE-Python package combined with the scikit-learn software. Hands on will be done in Python using Jupyter notebooks. No strong coding skills in Python are required.
Moritz Grosse-Wentrup
University of Vienna, Austria

Biography: Moritz Grosse-Wentrup is full professor and head of the Research Group Neuroinformatics at the University of Vienna, Austria. He develops machine learning algorithms that provide insights into how large-scale neural activity gives rise to (disorders of) cognition, and applies these algorithms in the domain of cognitive neural engineering, e.g., to build brain-computer interfaces for communication with severely paralyzed patients, design closed-loop neural interfaces for stroke rehabilitation, and develop personalized brain stimulation paradigms. He has received numerous awards for his work, including the 2011 Annual Brain-Computer Interface Research Award, the 2014 Teaching Award of the Graduate School of Neural Information Processing at the University of Tübingen, and the 2016 IEEE Brain Initiative Best Paper Award.
Lecture: Causal inference in neuroimaging
Abstract: Causal inference (CI) refers to the task of inferring causal relations from purely observational data. Causal inference is of particular interest in settings where ethical, financial, and logistic constraints limit interventional studies, e.g., in meuroimaging. In this lecture, I will introduce the mathematical foundations of causal reasoning and discuss various frameworks for causal inference in neuroimaging, including the potential outcomes framework, Causal Bayesian Networks, Granger causality, and Dynamic Causal Modelling. In the following Python tutorial, you will learn to apply encoding- and decoding models to carry out causal inference on simulated and actual neuroimaging data.
Georg Langs
Medical University of Vienna, Austria

Biography: Georg Langs studied Mathematics at Vienna University of Technology, and finished his PhD in Computer Vision at Vienna University of Technology and Graz University of Technology in 2007. He worked as a post-doctoral associate at the Applied Mathematics and Systems Laboratory at Ecole Centrale de Paris, and the GALEN Group at INRIA-Saclay, Ile de France with Nikos Paragios from 2007 to 2008. He was a Research Scientist at Computer Science and Artificial Intelligence Laboratory at Massachusetts Institute of Technology from 2009 to 2011, and joined the Faculty of Medical University of Vienna in 2011. He taught Computer Vision and Medical Imaging courses at Ecole Centrale de Paris, and teaches at Vienna University of Technology. He reviews for several Conferences and Journals, among them IEEE Transactions on Pattern Recogniton and Machine Intelligence, and IEEE Transactions on Medical Imaging. Georg Langs is the Head of the Computational Image Analysis and Radiology Lab (CIR) at the Medical University of Vienna.
Lecture: Magnetic Resonance Imaging Analysis via machine learning for structural and functional imaging
Máté Lengyel
University of Cambridge, UK & Central European University, Austria&Hungary

Biography: Máté Lengyel is Professor of Computational Neuroscience at the Department of Engineering, University of Cambridge, and a Senior Research Fellow at the Department of Cognitive Science, Central European University. Máté obtained an MSc in Cell, Development and Neurobiology and a PhD in Behavioural Neuroscience at Eötvös Loránd University, Budapest, Hungary, followed by a post-doctoral research fellowship at the Gatsby Computational Neuroscience Unit, UCL, and a visiting research fellowship at the Collegium Budapest Institute for Advanced Study. His interests span a broad range of levels of nervous system organisation, from sub-cellular and cellular through circuit and systems to behaviour and cognition. He studies these phenomena from computational, algorithmic/representational and neurobiological viewpoints. Computationally and algorithmically, he uses ideas from Bayesian approaches to statistical inference and reinforcement learning to characterise the goals and mechanisms of learning in terms of normative principles and behavioural results. He performs dynamical systems analyses of reduced biophysical models to understand the mapping of these mechanisms into cellular and network models. Máté collaborates very closely with experimental neuroscience groups, doing in vitro intracellular recordings, multi-unit recordings in behaving animals, and human psychophysical experiments.
Lecture: Bayesian inference in the brain and for the brain
Abstract: The Bayesian approach in statistics offers a principled and consistent mathematical calculus for inferring unknown quantities from known ones. This makes it a powerful theoretical framework for developing normative models of neural computation based on noisy, partial, or ambiguous inputs; in other words, in situations when the brain needs to act as a well-informed statistician — which is arguably almost always the case. Therefore, these models are relevant for a broad range of different levels of cognition, from perception, through causal reasoning, to motor control, and at different levels of brain organisation, from single cells, through networks and populations, to systems and behaviour. Naturally, the same mathematical-computational concepts and tools are also useful for us as researchers performing statistical analyses of our data. This tutorial will introduce the basic concepts of Bayesian inference, highlight some practical issues for consideration when applying them in practice, and demonstrate their usefulness through a series of case studies both in modelling and in data analysis.
Sebastian Tschiatschek
University of Vienna, Austria

Biography: Sebastian Tschiatschek is assistant professor of machine learning at the University of Vienna, Austria, since early 2020. Prior to that he spent a little more than two years at Microsoft Research in Cambridge, UK, where he was a senior researcher in the Machine Learning and Perception Group (now All Data AI). He obtained his PhD from Graz University of Technology, Austria, in 2014 and was a postdoctoral scholar at ETH Zurich from 2014 to 2017. His research interest includes probabilistic modeling, and exploration and imitation in reinforcement learning.
Lecture: Machine Learning for Neural Imaging
Abstract: In this session we study the typical machine learning pipeline and basic machine learning models for regression and classification, and briefly touch upon models for processing set-valued input data. We review applications of these models to selected problems in the medical domain and neural imaging. Finally, we conclude with a brief hands-on session in which we apply some of the studied models to standard machine learning datasets using an open-source framework.